Abstract
We examined the population structure of the introduced aphid, Myzus persicae collected mainly from its primary host, Prunus persica, in south-east Australia. Myzus persicae has been present in Australia since at least 1893. Samples were collected in the spring of 1998 from two mainland and three Tasmanian localities and isofemale lines were established in the laboratory. The reproductive mode (life cycle), karyotype and 17-locus microsatellite genotype of each clone were determined. All populations showed significant population differentiation (FST 0.058–0.202) even over small geographic distances (<50 km). All clones were karyotypically normal except for a subset of clones from one site that was exposed to the carbamate insecticide, Pirimor, the week prior to sampling. Those clones were heterozygous for an autosomal 1,3 translocation frequently associated in M. persicae with insecticide resistance. In contrast to other loci and despite being on different chromosomes, loci myz2A and M55A showed general and significant linkage disequilibrium. These loci may be affected by epistatic selection. We discuss the observed high clonal diversity, moderate but significant population differentiation, general conformance to Hardy-Weinberg equilibria and low linkage disequilibria with particular focus on the global population biology of M. persicae.
Similar content being viewed by others
Introduction
The geographic structure of a population is a function of the ecology and evolution of the organism and results from both demographic and genetic processes (Roderick, 1996). Demographic processes include rates of birth and death, immigration and emigration. Genetic processes include gene flow, mutation, genetic drift and natural selection. Gene flow, the movement of genes in and among populations, is a homogenising force counteracting the independent evolution of populations (Slatkin, 1987; Neigel, 1997).
For all organisms there is an interaction between demography and genetic structure (Roderick, 1996; Lacy, 1997) but aphids and other cyclical parthenogens have additional profound interactions associated with plasticity in reproductive mode. It is especially common for aphids that have spread to many parts of the world to exhibit a range of life cycle strategies (reproductive modes). Reproductive modes can encompass cyclical, functional or obligate parthenogenesis or a combination of all of these (Blackman, 1974). Additionally there is genetic variability in developmental regulation of reproductive mode within and among populations (Moran, 1992). Like most genetic variation, this is potentially visible to natural selection.
Myzus persicae, the peach-potato aphid, is a cosmopolitan, economically important pest aphid (Blackman and Eastop, 1984). It was first reported in Australia in 1910 in New South Wales (Zeck, 1928) but records in the Australian National Insect Collection indicate its presence since at least 1893 (from south-eastern Victoria, Record No.: AN20 4267 aphid database, M Carver, CSIRO Division of Entomology, Canberra, Australia). A vector of many plant viruses, it has developed resistance to insecticides throughout its geographic range (Devonshire et al, 1998). Organophosphate and carbamate resistance in M. persicae is based on amplification of a particular esterase gene (E4), and is associated with a translocation between autosomes 1 and 3 (Blackman et al, 1978; Devonshire, 1989). E4-based resistance is usually associated with obligately parthenogenetic lineages or male-producing obligate parthenogens (Spence and Blackman, 1998). Yet, whilst the translocation might be expected to cause problems at meiosis, it can pass through the sexual cycle (Blackman and Takada, 1977).
In this paper we investigate the population genetics of spring populations of M. persicae mainly from peach trees in Eastern Australia. Peach is the primary host of M. persicae, the plant on which the sexual phase occurs. Each egg hatching in spring on peach gives rise to a new clone. After several generations of parthenogenetic reproduction, these spring populations produce winged morphs that migrate to herbaceous plants where they continue parthenogenetic reproduction through the summer months, returning to peach in the autumn to reproduce sexually.
Materials and methods
Sample collection
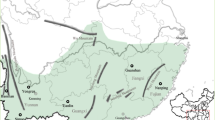
Collections were made from the primary host, Prunus persica, from one site in Bathurst, NSW, and from three sites in Tasmania (Figure 1) in the early spring (October) of 1998. Collectors were asked to cut twigs bearing aphids from the trees and send them to Macquarie University. Where possible a single clonal lineage was established from each twig. Later in the season (December) an additional collection was made from a vegetable garden in Canberra (Australian Capital Territory). Between eight and 19 isofemale lines were successfully established from each collection locality (full details in Table 1).
Map of Australia showing collection localities. Approximate geographic distances between Bathurst and Canberra = 155 km; Bathurst and the Tasmanian sites = 1080 km; Canberra and the Tasmanian sites = 860 km; Inverquarity and Sorell = 10 km; Inverquarity and New Town = 12.5 km; Sorell and New Town = 25 km.
Karyology
Chromosome preparations were made from somatic cells of young embryos dissected from fresh adult or fourth instar females, using the method of Blackman (1980). More than one embryo from an individual mother was macerated on a given slide. Several aphids per lineage were karyotyped. Unstained slides were examined using an Olympic Vanox microscope under phase contrast at 40× objective magnification. From three to 10 cells per slide were scored.
Characterisation of reproductive mode
Sexual morph production in aphids is largely a response to scotoperiod. In particular a critical length of dark- period but additionally a threshold low temperature must be reached before aphids will respond to night length (Lees, 1973; Blackman, 1975). An aphid clone can be maintained as a continuous parthenogenetic culture at long day and warm temperatures (eg, for M. persicae: 20°C with 16 h of light: 8 h of darkness). Sexual morph production, in clones capable of their production, is achieved experimentally by transferring adult parthenogenetic females to a short day, lower temperature environment. All our M. persicae cultures are continuously maintained on cabbage seedlings (var. Early Jersey Wakefield).
Three adult parthenogenetic females per cabbage seedling were transferred from 20°C 16L:8D to 15°C 10L:14D (short day). The aphids were allowed to reproduce at short day for 2 to 3 days after which time the adults were removed from the plants and destroyed. The young remaining on the plant (G1) were reared to adulthood and then transferred, three per plant, to new plants and allowed to reproduce for 2 to 3 days before serial transfer to new plants. In this way the offspring of G1 aphids were collected in sequential batches of G1 Batch 1 (G1B1), G1B2, G1B3 etc. When G1B1 reached adult moult they were transferred to a new plant, three aphids per plant. In most cyclically parthenogenetic (holocyclic) lineages reared under this regime, the G1B1 are alate parthenogenetic females (gynoparae) that produce mating females (oviparae). Males are commonly found in batches G1B5–G1B8. If a lineage produced both mating females and males it was recorded as cyclically parthenogenetic.
DNA extraction and amplification of microsatellite loci
DNA for microsatellite amplification was extracted by salting-out (Sunnucks and Hales, 1996). Microsatellite amplification was performed as described in Sloane et al (2001).
All isofemale lines were screened with 17 microsatellite loci: 10 cloned by ACCW from M. persicae (Sloane et al, 2001): M35A, M37A, M40A, M49A, M55A, M62A, M63A, M77, M86X and M107A, four cloned from M. persicae by Gavin Malarky (G Malarky, Natural History Museum, London, unpublished data): myz2A, myz3X, myz9A and myz25X, and the remaining three: S16bA, S17bX and S23A were cloned from Sitobion miscanthi (Wilson and Sunnucks, unpublished data). Information on chromosomal location is incorporated into the locus name of each spe- cies for which microsatellites have been cloned: A = autosomal, X = X-linked, and no superscript = currently unknown.
Population genetic analyses
Aphids bearing identical genotypes over all loci were identified using the program MSmacro (Park, 2000). Because inclusion of multiple clonal copies can lead to misinterpretation of population genetic analysis, datasets should be reduced to a single representative of each genotype provided that sufficient power exists in the genetic array (Sunnucks et al, 1997). Probabilities that a genotype might arise independently were approximated as the product of the local frequencies for the alleles in that genotype (Sunnucks et al, 1997). Accordingly, all population genetic analyses were performed on a data set that included only one copy of each clone at 17 loci, and none included the two clones with data missing from several loci (053S and 064NT).
Allele frequencies, Hardy-Weinberg equilibrium and linkage disequlibrium
Allele frequencies, mean number of alleles per locus (A), observed heterozygosity (Hobs), and an unbiased estimate expected under Hardy-Weinberg assumptions (HE, Nei, 1978) were calculated using BIOSYS-1 (Swofford and Selander, 1981). GENEPOP version 1.2 (Raymond and Rousset, 1995) was used to analyse Hardy-Weinberg equilibrium (HWE), linkage disequilibrium (LD) and to calculate FIS values (Weir and Cockerham, 1984). Population pairwise F-statistics for all population pairs and F-statistic confidence intervals based on bootstrap re-sampling were estimated with the FSTAT program (Weir and Cockerham, 1984; Goudet, 1995, 1999). Individuals possessing translocated karyotypes were not included in analysis of LD because the potentially different linkage of loci in normal and translocated individuals could confound estimation of LD. Sequential Bonferroni corrections were applied to multiple statistical tests (Rice, 1989; Sokal and Rohlf, 1995).
Estimating gene flow
Dispersal does not always result in gene flow and direct measurements of migration tend to underestimate rare but important gene flow (Slatkin, 1987). Because of the difficulties associated with directly measuring gene flow, indirect methods that use genetic data to infer movement can be used (Slatkin, 1985; Roderick, 1996).
Population pairwise genetic distances were calculated in Arlequin version 1.1 (Schneider et al, 1997). Pairwise FST values can be used as short-term genetic distances between populations (Reynolds et al, 1983; Slatkin, 1995). The null distribution of pairwise FST values under the hypothesis of no difference between populations is obtained by permuting haplotypes between populations. The P-value of the test is the proportion of 100 000 permutations leading to a FST value larger than or equal to the observed one.
Analysis of molecular variance (AMOVA)
The genetic structure of the data was investigated by AMOVA (Cockerham, 1969, 1973) using Arlequin Version 1.1 (Schneider et al, 1997). The approach is similar to other approaches based on analyses of variance of gene frequencies but also takes into account the number of mutations between molecular haplotypes (Excoffier et al, 1992). The significance of the fixation indices computed is tested using a non-parametric permutation approach (Excoffier et al, 1992). An AMOVA analysis was performed to test for genetic substructure among (i) all collections and (ii) the three geographically proximate Tasmanian collections.
Genetic distance analysis
All clones with missing data were eliminated from these analyses (019INV, 027B, 040B, 044S, 063NT) so n = 46. Individual pairwise and population pairwise genetic distances were calculated using Cavalli-Sforza and Edwards chord distance (C-S & E chord distance, Cavalli-Sforza and Edwards, 1967) because it has been shown, out of a range of common genetic distances, to produce the most reliable topologies with microsatellite loci (Takezaki and Nei, 1996). C-S & E chord distance transforms allele frequencies so as to standardise the distance with respect to genetic drift, and so reduces the impact of differences in starting allele frequencies (Swofford et al, 1996). Additionally, this measure is not heavily influenced by differences in within-taxon heterozygosity (Swofford et al, 1996). Raw microsatellite data were transformed into a pairwise genetic distance matrix using BIOSYS-1 (Swofford and Selander, 1981). The distance matrices generated were analysed using two-dimensional non-metric multidimensional scaling (NMDS) in SYN-TAX (Podani, 1995). NMDS is a form of ordination analysis that is useful for recovering non-hierarchical patterns of variation from overall estimates of genetic variation (Lessa, 1990). The application of non-hierarchical methods of analysis is particularly important when working within species because of the high probability of reticulate evolution (ie, within a species, because of gene flow, the relationship among populations is not likely to be hierarchical). NMDS does not assume linearity nor does it preclude the uncovering of hierarchical structure.
Results
Distribution of clones with translocated karyotypes
Only 7/55 (13%) of the karyotyped samples were found to carry the 1,3 autosomal translocation (Table 1). All seven samples were from the Inverquarity collection. These samples represent 70% of the lineages established from this site. The Inverquarity site had been sprayed with Pirimor insecticide in the week prior to sampling (M Williams, personal communication), in contrast to all other populations, which were known to be unsprayed.
Reproductive mode
The reproductive mode of 54 of the 57 isofemale lines established was characterised (three died out). All lines were cyclically parthenogenetic, indicating that those surviving the parthenogenetic phase would normally return to peach the following autumn to reproduce sexually.
Moderate genic and genotypic diversity
All loci except M77A and S23A were polymorphic in M. persicae (Table 2). Apart from the two monomorphic loci, and the highly polymorphic M49A locus (16 alleles), allelic diversity in the loci cloned from M. persicae was 3–9 alleles per locus (average = 6.3 alleles/locus). Allelic diversity at each of the two polymorphic loci cloned from Sitobion (S16bA and S17bX) was lower than that observed at the 13 polymorphic Myzus loci. Considering all loci, mean observed heterozygosity ranged from 0.00 to 1.00 per locus, and averaged 0.50–0.59 per population (Table 2 and 3).
Most genotypes (52 of 53) were collected only once. The one genotype sampled twice was 026BAT = 028BAT (identical at 34/34 alleles, and both were 2n = 12 non-translocated) (Table 1). Based on the observed allele frequencies in the Bathurst collection, the probability of independently generating the 026/028BAT genotype by sexual reproduction is 4.7 × 10−11. Thus 026BAT and 028BAT were assumed to be members of the same clone.
Small deviations from Hardy-Weinberg equilibrium and linkage disequilibrium
The Canberra and Inverquarity collections were in HWE at all polymorphic loci. Hardy-Weinberg exact tests in GENEPOP showed deviations from expectation: in the Bathurst collection at M62A, M63A, M107A and myz3X, in the Sorell collection at M55A and M62A and in the New Town collection at M62A, M86X and myz9A. Following Bonferroni correction within each sample, significant deviations from HWE remained in the Bathurst collection at loci M62A, M63A and myz3X and in the New Town collection at loci M62A, M86X and myz9. FIS values associated with all significant collection by locus deviations were significantly positive with the exception of M63A in the Bathurst collection which was significantly negative (FIS = −0.108, P = 0.0026) (Table 2).
No significant deviations from linkage equilibrium were observed within any population following Bonferroni correction (P > 0.05). However, locus pairs M55A/myz2A and M49A/M55A were in significant disequilibria (after Bonferroni correction, P < 0.05) when genotypes were pooled over populations.
Partitioning of genetic variation
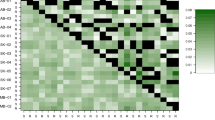
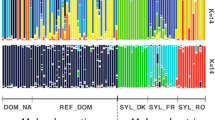
Most genetic variation was within collections (91.3%): among collection differentiation was modest but significant (8.7%, FST 0.087, P < 0.001). NMDS of the Cavalli-Sforza and Edwards chord distance was used to assess the genetic affinity of individual genotypes from the five collections (Figure 2). The far right of the cloud is composed of individuals from the Bathurst collection. Nested within Bathurst and towards the middle of the cloud are individuals from Canberra. Around the edge of the cloud, abutting Bathurst, and to the left and far left of the cloud are the three collections from Tasmania (all circular symbols, Figure 2). A similar analysis was performed to study among-collection structure. This analysis indicates that the Bathurst and Canberra collections are more closely related to each other than they are to the Tasmanian collections.
Similar patterns were seen at a local scale. Most of the genetic variation (92.6%) among the three geographically proximate Tasmanian collections was identified within collections and again moderate but significant genetic differentiation was identified between collections (FST = 0.074, P < 0.001).
Discussion
Colonization of Australia by Myzus persicae with at least moderate numbers of migrants
Hales et al (2000) characterised five Australian M. Persicae clones collected between 1990 and 1996, two from New South Wales, one from Queensland, one from Western Australia and one from Victoria. Over the 11 loci common to both, the 42 alleles identified in the earlier study were fully included in the 74 alleles discovered in this study. This large degree of overlap, especially in the light of low sample sizes scattered over sample dates and locations, leads us to believe that we have probably uncovered much of the allelic diversity at these loci for Australian M. persicae.
In a concurrent collaborative study, Terradot et al (1999) used nine microsatellite loci that were used in the present work (M35A, M37A, M40A, M63A, M77A, M107A, S16bA, S17bX, S23A), and identified 23 alleles in seven clones of European M. persicae. We found 20 of these alleles in our Australian collections. Whilst it is possible that some of these shared alleles could be the result of convergent evolution and not shared ancestry, it seems more likely that much of the allelic diversity observed in M. persicae in Australia was present in the colonists. Overall our data indicate that Australian M. persicae harbour a non-trivial proportion of the allelic diversity of the species in Europe, and are likely to be descended from a moderate or large number of colonists. In the absence of many more data from European aphids it is not possible to be more quantitative in this assessment, but clearly the situation is completely unlike that for Australian Sitobion aphids, of which only one or two successful clones per species seem to have colonised the continent (Sunnucks et al, 1996; Wilson et al, 1999).
Geographic structure of spring Myzus persicae populations in eastern Australia
All populations of Myzus persicae in Australia showed significant population differentiation even over small geographic (<50 km) distances (Table 4: FST 0.058–0.202). At first sight, this differentiation seems rather larger than that often reported for aphid populations from microsatellite data, such as the minimal differentiation observed over 60 km in S. avenae from southern England (Sunnucks et al, 1997), or that observed in S. avenae in France (Simon et al, 1999). However, much of the present differentiation may arise from stochastic effects including small sample sizes. There are several features characteristic of aphid populations that may explain the observed genetic differentiation in these M. persicae collections (Blackman, 1972, 1985). First, aphids are prone to major short-term changes in population size, which could have local stochastic or deterministic elements. Second, an aphid population is composed of a varying number of clones, represented at each point in time by an extremely variable number of identical individuals (barring mutation), with stochasticity due to egg hatching, clonal proliferation, immigration, local selection and extinction. Third, aphids are highly mobile and thus the structure of a population at any one time will depend on the number of immigrants (Blackman, 1972). Nonetheless, in addition to random factors some deterministic forces may also have contributed to population differentiation, as explored in the following sections.
Linkage disequilibria: physical linkage, genetic stochasticity or selection?
The loci used in this study are known to represent four autosomal and one X-linked linkage groups: (i) myz2A – myz9A – S16bA, (ii) M49A – M55A – M37A – M62A, (iii) M35A – M63A – M107A, (iv) M40A and loci myz3X – myz25X – M86X – S17bX are X-linked (Sloane et al, 2001). Two pairs of loci, M49A/M55A (same chromosome) and M55A/myz2A (not same chromosome), were found to be in significant linkage disequilibrium over all genotypes. Linkage disequilibrium between loci M49A and M55A may be caused by physical linkage and population contractions. However, as M55A and myz2A are in different linkage groups yet show general LD over all genotypes, their association may result from selective epistasis.
Association of the heterozygous 1,3 autosomal translocation with the presence of insecticide
Translocated clones were identified only in the collection from Inverquarity, Tasmania, from a nectarine orchard that had been sprayed with the insecticide Pirimor in the week prior to sampling. Insecticide treatment has selected for resistant clones bearing the translocation (Blackman et al, 1978; Devonshire et al, 1998). From their presence in this but not other collections, we infer a history of insecticide use in this orchard.
Genetic diversity and reproductive mode
The application of microsatellites to populations of aphids has overturned several classical predictions about patterns of genotypic diversity and heterozygosity. For example, Sitobion avenae in England was thought to be largely obligately parthenogenetic and therefore would have been predicted to be represented by few clones, yet most individuals sampled bear a unique microsatellite genotype (Sunnucks et al, 1997). Also counter to general expectation (eg Amos, 1999), S. avenae in France with frequent sexual reproduction have significantly lower heterozygosity than do obligately parthenogenetic lines, and indeed the most regularly sexual lineages show significant homozygous excess (Simon et al, 1999).
The present results extend observations of genetic diversity in cyclic parthenogens. Based on results from S. avenae in France (Simon et al, 1999), data from cyclically parthenogenetic (holocyclic) M. persicae in Australia might have been expected to show strong positive FIS at many loci (FIS: is a measure of the deviation from Hardy-Weinberg proportions within subpopulations; a positive FIS indicates a deficiency of heterozygotes where a negative FIS indicates an excess of heterozygotes). However, such a general pattern was not observed. Three of the five populations analysed here showed more negative than positive FIS, and two populations with more loci showing positive than negative FIS still had five loci with negative FIS. Further, at least some of the positive FIS values may be the result of null (non-amplifying) alleles at a microsatellite locus (eg, locus M62A showed significant deviations from HWE in three of the five collections, and FIS values are strongly positive in all but the Canberra collection). Thus the finding that French S. avenae with a regular sexual cycle are significantly and generally homozygous at microsatellite loci is not parallelled in Australian M. persicae with a regular sexual cycle: these latter aphids are generally quite close to genetic equilibrium. This suggests that the observations in French Sitobion may not represent a general phenomenon, but one worthy of further investigation.
Previous population studies in Myzus persicae
Like many aphids, population studies of M. persicae using allozyme markers have found little to no genetic variation (May and Holbrook, 1978; Wool et al, 1978; Brookes and Loxdale, 1987; Hales et al, 1997). In fact, in M. persicae, the only loci that have been found to be polymorphic are the esterase loci. Esterase polymorphisms in M. persicae have been strongly selected as a result of their association with organophosphate resistance and with cross-resistance to other insecticide groups including carbamates (Devonshire, 1989). This lack of genetic variation at allozyme loci is not fully understood particularly because it is not generally mirrored by other molecular markers which have frequently revealed high levels of genic and genotypic diversity. A random amplified fragment length polymorphism (RAPD-PCR) study of M. persicae in 74 isofemale lines collected in Spain revealed that almost every isofemale line was different (Martinez-Torres et al, 1997). A similar level of genetic variation was found using an rDNA fingerprint technique on Scottish populations of M. persicae (Fenton et al, 1998). In that study 80 fingerprints amongst 276 isofemale lines were identified. Thirty percent of the isofemale lines were found to have the same fingerprint indicating the presence of a single widespread clone in Scotland that was found to be present in samples over a 4-year period (1992–1996). In addition to this common clone other genotypes were sampled over 2 years suggesting that local over-wintering of asexual clones is the most common source of spring M. persicae populations in Scotland.
Microsatellite analysis of south-eastern Australian M. persicae has revealed high levels of genic and genotypic diversity and is consistent with the levels of genotypic diversity detected in Spanish and Scottish M. persicae populations using multilocus DNA markers (Martinez-Torres et al, 1997; Fenton et al, 1998). Unlike in Scotland, where peach is rare, no common and widespread clone was identified in these samples from south-eastern Australia. This, in large part, is attributable to the fact that most of our collections were made from peach trees in spring, and were offspring of the founding females that emerged from sexually-produced eggs. Studies of M. persicae populations in Western Australia, where M. persicae is believed to over-winter exclusively by parthenogenesis on secondary host plants (Blackman, 1974), may find a pattern similar to that described for M. persicae in Scotland (Fenton et al, 1998) and S. avenae in eastern France (Haack et al, 2000).
Conclusions
Unlike several aphid species studied with microsatellites, M. persicae in Australia are generally close to genetic equilibrium. The species has maintained regular sexual reproduction following migration to lower latitudes, at least in areas with cold winters, and has colonised in sufficient numbers to retain an appreciable proportion of allelic diversity. This M. persicae dataset also represents the second of two counter-examples of the intriguing result that French S. avenae with an obligate sexual cycle have significant heterozygous deficits.
References
Amos, W (1999). A comparative approach to the study of microsatellite evolution. In: Goldstein DB, Schlötterer C (eds) A comparative approach to the study of microsatellite evolution, Oxford University Press: Oxford, pp 66–79.
Blackman, RL (1972). The inheritance of life-cycle differences in Myzus persicae (Sulz.) (Hem., Aphididae). Bull Entomolog Res, 62: 281–294.
Blackman, RL (1974). Life-cycle variation of Myzus persicae (Sulz.) (Hom., Aphididae) in different parts of the world, in relation to genotype and environment. Bull Entomolog Res, 63: 595–607.
Blackman, RL (1975). Photoperiodic determination of the male and female sexual morphs of Myzus persicae. J Insect Physiol, 21: 435–453.
Blackman, RL (1980). Chromosome numbers in the Aphididae and their taxonomic significance. Systemat Entomol, 5: 7–25.
Blackman, RL (1985). Aphid cytology and genetics. In: Aphid Cytology and Genetics, Wydawnictowo Polsk Akad: Nauk Warsaw, Jablonna, pp 171–237.
Blackman, RL, Takada, H (1977). The inheritance of natural chromosomal polymorphisms in the aphid Myzus persicae (Sulzer). Genetica, 47: 9–15.
Blackman, RL, Takada, H, Kawakami, K (1978). Chromosomal rearrangement involved in insecticide resistance of Myzus persicae. Nature, 271: 450–452.
Blackman, RL, Eastop, VF (1984). Aphids on the World's Crops: an identification and information guide. John Wiley and Sons: Chichester.
Brookes, CP, Loxdale, HD (1987). Survey of enzyme variation in British populations of Myzus persicae (Sulzer) (Hemiptera: Aphididae) on crops and weed hosts. Bull Entomolog Res, 77: 83–99.
Cavalli-Sforza, LL, Edwards, AWF (1967). Phylogenetic analysis: models and estimation procedures. Evolution, 21: 550–570.
Cockerham, CC (1969). Variance of gene frequencies. Evolution, 23: 72–84.
Cockerham, CC (1973). Analyses of gene frequencies. Genetics, 74: 679–700.
Devonshire, AL (1989). Resistance of aphids to insecticides. In: Minks AK, Harrewijn P (eds) Resistance of Aphids to Insecticides, Elsevier: Amsterdam, pp 123–139.
Devonshire, AL, Field, LM, Foster, SP, Moores, GD, Williamson, MS, Blackman, RL (1998). The evolution of insecticide resistance in the peach-potato aphid, Myzus persicae. Phil Trans Royal Soc Lond Ser B, 353: 1677–1684.
Excoffier, L, Smouse, PE, Quattro, JM (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics, 131: 479–491.
Fenton, B, Woodford, JAT, Malloch, G (1998). Analysis of clonal diversity of the peach-potato aphid, Myzus persicae (Sulzer), in Scotland, UK and evidence for the existence of a predominant clone. Mol Ecol, 7: 1475–1487.
Goudet, J (1995). FSTAT (version 1.2) A computer program to calculate F-statistics. J Hered, 86: 485–486.
Goudet, J (1999). FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.8). Updated from Goudet (1995).
Haack, L, Simon, J-C, Gauthier, J-P, Plantegenest, M, Dedryver, C-A (2000). Predominance of generalist clones in a cyclically parthenogenetic organism evidenced by combined demographic and genetic analyses. Mol Ecol, 9: 2055–2066.
Hales, D, Wilson, ACC, Spence, JM, Blackman, RL (2000). Confirmation that Myzus antirrhinii (Macchiati) (Hemiptera: Aphididae) occurs in Australia, using morphometrics, microsatellite typing, and analysis of novel karyotypes by fluorescence in situ hybridisation. Aust J Entomol, 39: 123–129.
Hales, DF, Tomiuk, J, Wöhrmann, K, Sunnucks, P (1997). Evolutionary and genetic aspects of aphid biology: a review. Euro J Entomol, 94: 1–55.
Lacy, RC (1997). Importance of genetic variation to the viability of mammalian populations. J Mammal, 78: 320–335.
Lees, AD (1973). Photoperiodic time measurement in the aphid Megoura viciae. J Insect Physiol, 19: 2279–2316.
Lessa, EP (1990). Multidimensional analysis of geographic genetic structure. Syst Zool, 39: 242–252.
Martinez-Torres, D, Carrio, R, Latorre, A, Simon, JC, Hermoso, A, Moya, A (1997). Assessing the nucleotide diversity of three aphid species by RAPD. J Evolu Biol, 10: 459–477.
May, B, Holbrook, FR (1978). Absence of genetic variability in the green peach aphid Myzus persicae (Hemiptera: Aphididae). Annals Entomol Soc Am, 71: 809–812.
Moran, NA (1992). The evolution of aphid lifecycles. Ann Rev Entomol, 37: 321–348.
Nei, M (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: 583–590.
Neigel, JE (1997). A comparison of alternative strategies for estimating gene flow from genetic markers. Ann Rev Ecol Systemat, 28: 105–128.
Park, S (2000). Autosomal Microsatellite Data Macros for Excel:Freeware excel macros for checking, formatting and analysing autosomal microsatellite data. http://acer.gen.tcd.ie/~sdepark/MSmacros.htm, Dublin.
Podani, J (1995). SYN-TAX 502Mac: Computer programs for multivariate data analysis on the Macintosh system. Scientia Publishing: Budapest.
Raymond, M, Rousset, F (1995). GENEPOP (version 1.2): a population genetics software for exact tests and ecumenicism. J Hered, 86: 248–249.
Reynolds, J, Weir, BS, Cockerham, CC (1983). Estimation of the coancestry coefficient: basis for a short-term genetic distance. Genetics, 105: 767–779.
Rice, WR (1989). Analysing tables of statistical tests. Evolution, 43: 223–225.
Roderick, GK (1996). Geographic structure of insect populations: gene flow, phylogeography, and their uses. Ann Rev Entomol, 41: 325–352.
Schneider, S, Kueffer, J-M, Roessli, D, Excoffier, L (1997). Arelquin ver. 1.1: A software for population genetic data analysis, Version 1.1, Genetics and Biometry Laboratory: University of Geneva, Switzerland.
Simon, JC, Baumann, S, Sunnucks, P, Hebert, PDN, Pierre, J-S et al (1999). Reproductive mode and population genetic structure of the cereal aphid Sitobion avenae studied using phenotypic and microsatellite markers. Mol Ecol, 8: 531–545.
Slatkin, M (1985). Gene flow in natural populations. Ann Rev Ecol Systemat, 16: 393–430.
Slatkin, M (1987). Gene flow and the geographic structure of natural populations. Science, 236: 87–92.
Slatkin, M (1995). A measure of population subdivision based on microsatellite allele frequencies. Genetics, 139: 457–462.
Sloane, MA, Sunnucks, P, Wilson, ACC, Hales, DF (2001). Microsatellite isolation, linkage group identification and determination of recombination frequency in the peach-potato aphid, Myzus persicae (Sulzer) (Hemiptera: Aphididae). Genetic Res Cambridge, 77: 251–260.
Sokal, RR, Rohlf, FJ (1995). Biometry. WH Freeman & Co: San Francisco.
Spence, JM, Blackman, RL (1998). Chromosomal rearrangements in the Myzus persicae group and their evolutionary significance. In: Nieto Nafría JM, Dixon AFG (eds) Chromosomal Rearrangements in the Myzus persicae Group and Their Evolutionary Significance, Universidad De León Secretariado de Publicacions: León (Spain), pp 113–118
Sunnucks, P, de Barro, PJ, Lushai, G, Maclean, N, Hales, DF (1997). Genetic structure of an aphid studied using microsatellites: cyclic parthenogenesis, differentiated lineages, and host specialization. Mol Ecol, 6: 1059–1073.
Sunnucks, P, England, PE, Taylor, AC, Hales, DF (1996). Micro-satellite and chromosome evolution of parthenogenetic Sitobion aphids in Australia. Genetics, 144: 747–756.
Sunnucks, P, Hales, DF (1996). Numerous transposed sequences of mitochondrial cytochrome oxidase I–II in aphids of the genus Sitobion (Hemiptera: Aphididae). Molec Biol Evol, 13: 510–523.
Swofford, DL, Selander, RB (1981). BIOSYS-1: A FORTRAN program for the comprehensive analysis of electrophoretic data in population genetics and systematics. J Hered, 72: 281–283.
Swofford, DL, Olsen, GJ, Waddell, PJ, Hillis, DM (1996). Phylogenetic Inference. In: Hillis DM, Moritz C, Mable BK (eds) Molecular Systematics, Second, Sinauer Associates: Sunderland,Massachusetts, USA, pp 407–514.
Takezaki, N, Nei, M (1996). Genetic distances and reconstructing of phylogenetic trees from microsatellite DNA. Genetics, 144: 389–399.
Terradot, L, Simon, J-C, Leterme, N, Bourdin, D, Wilson, ACC, Gauthier, J-P, Robert, Y (1999). Molecular characterization of clones of the Myzus persicae complex (Hemiptera: Aphididae) differing in their ability to transmit the potato leafroll luteovirus (PLRV). Bull Entomolog Res, 89: 355–363.
Weir, BS, Cockerham, CC (1984). Estimating F-statistics for the analysis of population structure. Evolution, 38: 1358–1370.
Wilson, ACC, Sunnucks, P, Hales, DFH (1999). Microevolution, low clonal diversity and genetic affinities of parthenogenetic Sitobion aphids in New Zealand. Molec Ecol, 8: 1655–1666.
Wool, D, Bunting, S, van Emden, HF (1978). Electrophoretic study of genetic variation in British Myzus persicae (Sulz.) (Hemiptera, Aphididae). Biochem Genet, 16: 987–1003.
Zeck, EH (1928). The green peach aphid, Myzus persicae (Sulzer). Agricult Gazette N South Wales, 39: 147–154.
Acknowledgements
Gavin Malarky, then at the Natural History Museum, London, cloned four of the microsatellite loci used in this study (myz2, myz3, myz9 and myz25). Many thanks to Tony Hellier (Bathurst) and Margaret Williams (Hobart) for collecting aphids from peach trees and to Mat Sloane for technical assistance. The work was supported by an Australian Research Council Grant to DFH & PS. Support during writing to ACCW came from National Science Foundation grants to RK Grosberg. This paper is Key Centre for Biodiversity & Bioresources publication no. 353.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wilson, A., Sunnucks, P., Blackman, R. et al. Microsatellite variation in cyclically parthenogenetic populations of Myzus persicae in south-eastern Australia. Heredity 88, 258–266 (2002). https://doi.org/10.1038/sj.hdy.6800037
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800037
Keywords
This article is cited by
-
Tracking the global dispersal of a cosmopolitan insect pest, the peach potato aphid
BMC Ecology (2009)
-
Does insecticide resistance alone account for the low genetic variability of asexually reproducing populations of the peach-potato aphid Myzus persicae?
Heredity (2005)
-
Fine-scale genetic structure of grape phylloxera from the roots and leaves of Vitis
Heredity (2004)
-
Spatial and temporal genetic variability in French populations of the peach–potato aphid, Myzus persicae
Heredity (2003)
-
Population genetic structure of the lettuce root aphid, Pemphigus bursarius (L.), in relation to geographic distance, gene flow and host plant usage
Heredity (2003)